Abstract
Here, in a multi-ancestry genome-wide association study meta-analysis of kidney cancer (29,020 cases and 835,670 controls), we identified 63 susceptibility regions (50 novel) containing 108 independent risk loci. In analyses stratified by subtype, 52 regions (78 loci) were associated with clear cell renal cell carcinoma (RCC) and 6 regions (7 loci) with papillary RCC. Notably, we report a variant common in African ancestry individuals (rs7629500) in the 3′ untranslated region of VHL, nearly tripling clear cell RCC risk (odds ratio 2.72, 95% confidence interval 2.23–3.30). In cis-expression quantitative trait locus analyses, 48 variants from 34 regions point toward 83 candidate genes. Enrichment of hypoxia-inducible factor-binding sites underscores the importance of hypoxia-related mechanisms in kidney cancer. Our results advance understanding of the genetic architecture of kidney cancer, provide clues for functional investigation and enable generation of a validated polygenic risk score with an estimated area under the curve of 0.65 (0.74 including risk factors) among European ancestry individuals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All GWAS summary statistics are available on dbGaP (phs003505.v1.p1) and GWAS Catalog (GCST90320043–GCST90320065). Individual-level data from the new NCI-3 scan are available on dbGaP (phs003505.v1.p1). Data from previously published scans are also available on dbGaP (NCI-1, phs000351.v1.p1; NCI-2, phs001736.v2.p1; USKC, phs000863.v1.p1; IARC-2, phs001271.v1.p1; MDA and MDA-OncoArray, phs003505.v1.p1). The individual-level data from the IARC-1 and UK scans have not been deposited in dbGaP or any other data archive site given decisions by the institutional ethics review boards for these projects. The data from these scans are available upon reasonable request through internal processes unique to each institution. Such requests can be made in writing to the principal investigators (IARC: P. Brennan, pbrennan@iarc.fr and UK: R.H., richard.houlston@icr.ac.uk); the time frame from request to receipt of data is approximately 4–6 weeks. The UK Biobank analysis was conducted via application number 86140 (https://www.ukbiobank.ac.uk/). The Finnish biobank data included in FinnGen can be accessed via Fingenious services at https://site.fingenious.fi/en/ (ref. 80) managed by FINBB. Finnish Health register data can be applied for via Findata at https://findata.fi/en/data/ (ref. 81). The full GWAS results of the BBJ are available via the website of the Japanese ENcyclopedia of GEnetic Associations by Riken (JENGER) at http://jenger.riken.jp/en/ (ref. 82; case–control GWAS no. 156). Function annotation enrichment was performed with the annotation data provided via the GARFIELD package at https://www.ebi.ac.uk/birney-srv/GARFIELD/ (ref. 83). Position weight matrices for transcription factor-binding sites as cataloged in HOCOMOCO were provided along with the motifbreakR R package, in the associated MotifDb database. ChIP-Seq data reported by Schmid et al.18 are publicly available through the Gene Expression Omnibus (GEO) database under the accession codes: GSE120885 (HIF-1α, HIF-2α and HIF-1β ChIP-seq in RCC4 cells) and GSE67237 (HIF-2α and HIF-1β ChIP-seq in 786-O cells). Epigenomic charting data (H3K27ac peaks) generated by Nassar et al.37 are publicly available through GEO database under accession code GSE188486; the sample attributes are mentioned in Supplementary Table 1 of the corresponding paper. GTEx v8 and TCGA data can be accessed via GTEx and Genomic Data Commons at https://gtexportal.org/home/ (ref. 84) and https://portal.gdc.cancer.gov/repository (ref. 85), respectively. Additionally, eQTLs for TCGA were queried via the PancanQTL database at http://gong_lab.hzau.edu.cn/PancanQTL/ (ref. 86).
Code availability
Code used in performing the liftover of summary statistics and fixed-effects GWAS meta-analyses (version 2022-12-23) is available via GitHub at https://github.com/freeseek/score (ref. 72). No previously unreported custom computer code or algorithm was used to generate results.
References
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Znaor, A., Lortet-Tieulent, J., Laversanne, M., Jemal, A. & Bray, F. International variations and trends in renal cell carcinoma incidence and mortality. Eur. Urol. 67, 519–530 (2015).
Cancer stat facts: kidney and renal pelvis cancer. National Cancer Institute https://seer.cancer.gov/statfacts/html/kidrp.html (2023).
Cancer Facts & Figures 2022. American Cancer Society https://www.cancer.org/research/cancer-facts-statistics/all-cancer-facts-figures/cancer-facts-figures-2022.html (2022).
Chow, W. H., Scelo, G. & Tarone, R. E. in Schottenfeld and Fraumeni Cancer Epidemiology and Prevention 4th edn (eds Thun, M. J. et al.) Ch. 51, 961–976 (Oxford Univ. Press, 2018).
Lopez-Beltran, A. et al. 2009 update on the classification of renal epithelial tumors in adults. Int. J. Urol. 16, 432–443 (2009).
Haas, N. B. & Nathanson, K. L. Hereditary kidney cancer syndromes. Adv. Chronic Kidney Dis. 21, 81–90 (2014).
Andreou, A. et al. Elongin C (ELOC/TCEB1)-associated von Hippel-Lindau disease. Hum. Mol. Genet. 31, 2728–2737 (2022).
Lang, M. et al. Clinical and molecular characterization of microphthalmia-associated transcription factor (MITF)-related renal cell carcinoma. Urology 149, 89–97 (2021).
Schmidt, L. S. et al. PRDM10 RCC: a Birt–Hogg–Dube-like syndrome associated with lipoma and a highly penetrant, aggressive renal tumors morphologically resembling type 2 papillary renal cell carcinoma. Urology 179, 58–70 (2023).
Scelo, G. et al. Genome-wide association study identifies multiple risk loci for renal cell carcinoma. Nat. Commun. 8, 15724 (2017).
Grampp, S. et al. Genetic variation at the 8q24.21 renal cancer susceptibility locus affects HIF binding to a MYC enhancer. Nat. Commun. 7, 13183 (2016).
Schodel, J. et al. Common genetic variants at the 11q13.3 renal cancer susceptibility locus influence binding of HIF to an enhancer of cyclin D1 expression. Nat. Genet. 44, 420–425 (2012).
Bigot, P. et al. Functional characterization of the 12p12.1 renal cancer-susceptibility locus implicates BHLHE41. Nat. Commun. 7, 12098 (2016).
Colli, L. M. et al. Altered regulation of DPF3, a member of the SWI/SNF complexes, underlies the 14q24 renal cancer susceptibility locus. Am. J. Hum. Genet. 108, 1590–1610 (2021).
Riscal, R. et al. Cholesterol auxotrophy as a targetable vulnerability in clear cell renal cell carcinoma. Cancer Discov. 11, 3106–3125 (2021).
Grampp, S. et al. Multiple renal cancer susceptibility polymorphisms modulate the HIF pathway. PLoS Genet. 13, e1006872 (2017).
Schmid, V. et al. Co-incidence of RCC-susceptibility polymorphisms with HIF cis-acting sequences supports a pathway tuning model of cancer. Sci. Rep. 9, 18768 (2019).
Patel, S. A. et al. The renal lineage factor PAX8 controls oncogenic signalling in kidney cancer. Nature 606, 999–1006 (2022).
Han, B. & Eskin, E. Random-effects model aimed at discovering associations in meta-analysis of genome-wide association studies. Am. J. Hum. Genet. 88, 586–598 (2011).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Cancer Genome Atlas Research Network et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Uhlen, M. et al. A pathology atlas of the human cancer transcriptome. Science 357, eaan2507 (2017).
Singhal, S. S., Yadav, S., Drake, K., Singhal, J. & Awasthi, S. Hsf-1 and POB1 induce drug sensitivity and apoptosis by inhibiting Ralbp1. J. Biol. Chem. 283, 19714–19729 (2008).
Oosterhoff, J. K., Kuhne, L. C., Grootegoed, J. A. & Blok, L. J. EGF signalling in prostate cancer cell lines is inhibited by a high expression level of the endocytosis protein REPS2. Int. J. Cancer 113, 561–567 (2005).
Oosterhoff, J. K., Penninkhof, F., Brinkmann, A. O., Anton Grootegoed, J. & Blok, L. J. REPS2/POB1 is downregulated during human prostate cancer progression and inhibits growth factor signalling in prostate cancer cells. Oncogene 22, 2920–2925 (2003).
Zhang, H., Duan, C. J., Zhang, H., Cheng, Y. D. & Zhang, C. F. Expression and clinical significance of REPS2 in human esophageal squamous cell carcinoma. Asian Pac. J. Cancer Prev. 14, 2851–2857 (2013).
He, X. Y. et al. Liver X receptor agonists exert antitumor effects against hepatocellular carcinoma via inducing REPS2 expression. Acta Pharmacol. Sin. 44, 635–646 (2023).
Du, J. et al. Cytoplasmic localization of IRF5 induces Wnt5a/E-cadherin degradation and promotes gastric cancer cells metastasis. Cancer Gene Ther. 30, 866–877 (2023).
Bi, X. et al. Loss of interferon regulatory factor 5 (IRF5) expression in human ductal carcinoma correlates with disease stage and contributes to metastasis. Breast Cancer Res 13, R111 (2011).
Massimino, M. et al. IRF5 promotes the proliferation of human thyroid cancer cells. Mol. Cancer 11, 21 (2012).
Guiteras, J. et al. The gene silencing of IRF5 and BLYSS effectively modulates the outcome of experimental lupus nephritis. Mol. Ther. Nucleic Acids 24, 807–821 (2021).
Zou, Y., Carbonetto, P., Wang, G. & Stephens, M. Fine-mapping from summary data with the ‘Sum of Single Effects’ model. PLoS Genet. 18, e1010299 (2022).
Kichaev, G. et al. Integrating functional data to prioritize causal variants in statistical fine-mapping studies. PLoS Genet. 10, e1004722 (2014).
Iotchkova, V. et al. GARFIELD classifies disease-relevant genomic features through integration of functional annotations with association signals. Nat. Genet. 51, 343–353 (2019).
Kulakovskiy, I. V. et al. HOCOMOCO: towards a complete collection of transcription factor binding models for human and mouse via large-scale ChIP-Seq analysis. Nucleic Acids Res. 46, D252–D259 (2018).
Nassar, A. H. et al. Epigenomic charting and functional annotation of risk loci in renal cell carcinoma. Nat. Commun. 14, 346 (2023).
Mucci, L. A. et al. Familial risk and heritability of cancer among twins in nordic countries. JAMA 315, 68–76 (2016).
Zhang, Y., Qi, G., Park, J. H. & Chatterjee, N. Estimation of complex effect-size distributions using summary-level statistics from genome-wide association studies across 32 complex traits. Nat. Genet. 50, 1318–1326 (2018).
Zhang, Y. D. et al. Assessment of polygenic architecture and risk prediction based on common variants across fourteen cancers. Nat. Commun. 11, 3353 (2020).
Linehan, W. M., Srinivasan, R. & Schmidt, L. S. The genetic basis of kidney cancer: a metabolic disease. Nat. Rev. Urol. 7, 277–285 (2010).
Chan, J. J., Tabatabaeian, H. & Tay, Y. 3′UTR heterogeneity and cancer progression. Trends Cell Biol. 33, 568–582 (2023).
Yang, Y. et al. The deubiquitinase USP38 promotes NHEJ repair through regulation of HDAC1 activity and regulates cancer cell response to genotoxic insults. Cancer Res. 80, 719–731 (2020).
Cancer Genome Atlas Research Network et al. Comprehensive molecular characterization of papillary renal-cell carcinoma. N. Engl. J. Med. 374, 135–145 (2016).
Olshan, A. F. et al. Racial difference in histologic subtype of renal cell carcinoma. Cancer Med 2, 744–749 (2013).
Usher-Smith, J., Simmons, R. K., Rossi, S. H. & Stewart, G. D. Current evidence on screening for renal cancer. Nat. Rev. Urol. 17, 637–642 (2020).
Jin, Y., Schaffer, A. A., Feolo, M., Holmes, J. B. & Kattman, B. L. GRAF-pop: a fast distance-based method to infer subject ancestry from multiple genotype datasets without principal components analysis. G3 9, 2447–2461 (2019).
Database of Genotypes and Phenotypes (NCBI, 2014); https://www.ncbi.nlm.nih.gov/gap/
Brown, D. W., Myers, T. A. & Machiela, M. J. PCAmatchR: a flexible R package for optimal case-control matching using weighted principal components. Bioinformatics 37, 1178–1181 (2021).
Chen, S. et al. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 625, 92–100 (2022).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 590, 290–299 (2021).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
Purdue, M. P. et al. Genome-wide association study of renal cell carcinoma identifies two susceptibility loci on 2p21 and 11q13.3. Nat. Genet. 43, 60–65 (2011).
Wu, X. et al. A genome-wide association study identifies a novel susceptibility locus for renal cell carcinoma on 12p11.23. Hum. Mol. Genet 21, 456–462 (2012).
Shu, X. et al. Potential susceptibility loci identified for renal cell carcinoma by targeting obesity-related genes. Cancer Epidemiol. Biomark. Prev. 26, 1436–1442 (2017).
Henrion, M. et al. Common variation at 2q22.3 (ZEB2) influences the risk of renal cancer. Hum. Mol. Genet 22, 825–831 (2013).
Purdue, M. P. et al. A genome-wide association study of renal cell carcinoma among African Americans. Cancer Epidemiol. Biomark. Prev. 23, 209–214 (2014).
Manichaikul, A. et al. Robust relationship inference in genome-wide association studies. Bioinformatics 26, 2867–2873 (2010).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
Nagai, A. et al. Overview of the BioBank Japan Project: study design and profile. J. Epidemiol. 27, S2–S8 (2017).
Index (BioBank Japan, 2017); https://biobankjp.org/en/index.html
Terao, C. et al. Chromosomal alterations among age-related haematopoietic clones in Japan. Nature 584, 130–135 (2020).
Tanaka, N. et al. Eight novel susceptibility loci and putative causal variants in atopic dermatitis. J. Allergy Clin. Immunol. 148, 1293–1306 (2021).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Zhou, W. et al. Efficiently controlling for case-control imbalance and sample relatedness in large-scale genetic association studies. Nat. Genet. 50, 1335–1341 (2018).
Begg, C. B. & Zhang, Z. F. Statistical analysis of molecular epidemiology studies employing case-series. Cancer Epidemiol. Biomark. Prev. 3, 173–175 (1994).
Lyon, M. S. et al. The variant call format provides efficient and robust storage of GWAS summary statistics. Genome Biol. 22, 32 (2021).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
freeseek (Github, 2022); https://github.com/freeseek/score
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Lin, S. H., Brown, D. W. & Machiela, M. J. LDtrait: an online tool for identifying published phenotype associations in linkage disequilibrium. Cancer Res. 80, 3443–3446 (2020).
Consortium, E. P. et al. Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 583, 699–710 (2020).
Coetzee, S. G., Coetzee, G. A. & Hazelett, D. J. motifbreakR: an R/Bioconductor package for predicting variant effects at transcription factor binding sites. Bioinformatics 31, 3847–3849 (2015).
Yang, J. et al. Genetic variance estimation with imputed variants finds negligible missing heritability for human height and body mass index. Nat. Genet. 47, 1114–1120 (2015).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Fingenius (FINBB, 2024); https://site.fingenious.fi/en/
Data (FINDATA, 2017); https://findata.fi/en/data/
Laboratory for Statistical and Translational Genetics (Japanese ENcyclopedia of GEnetic associations by Riken, 2021); http://jenger.riken.jp/en/
GARFIELD (EMBL-EBI, 2015); https://www.ebi.ac.uk/birney-srv/GARFIELD/
GTEx Portal (GTEx, 2017); https://gtexportal.org/home/
Repository (Genomic Data Commons, 2024); https://portal.gdc.cancer.gov/repository
Cis-eQTLs and Trans-eQTLs in 33 Cancer Types (PancanQTL, 2018); http://gong_lab.hzau.edu.cn/PancanQTL/
Acknowledgements
We thank O. Jahagirdar and A. Klein for their efforts in implementing various computational pipelines used in this project. This research was supported by the Intramural Research Program of the National Cancer Institute, National Institutes of Health, US Department of Health and Human Services, as well as with Federal funds from the National Cancer Institute under contract no. 75N91019D00024. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the US Government. Where authors are identified as personnel of the International Agency for Research on Cancer/World Health Organization, the authors alone are responsible for the views expressed in this article and they do not necessarily represent the decisions, policy or views of the International Agency for Research on Cancer/World Health Organization. Acknowledgments and funding sources for participating centers are provided in Supplementary Table 17.
Author information
Authors and Affiliations
Consortia
Contributions
M.P.P., D.D., M.J.M., B.R.G., P. Brennan, J.G. and S.J.C. contributed to the design and execution of the overall study. M.Y., N.R.C., B.D.H., M.R.M. and A.A.H. performed the experiments. M.P.P., D.D., M.J.M., B.R.G., T.W., D.O., S.C., A. Liu, K.H., K.M.B., J.R. and S.J.C. contributed to the design and execution of the statistical analysis. M.P.P., D.D., M.J.M., B.R.G. and S.J.C. contributed to the first draft of the paper. P. Brennan, J.G. and S.J.C. edited the paper. A.F.-I., P.S., A. Liu, C.W., S.O.A., J. Larkin, S.C.Z., M.S., K.H., A.H., K.A.L., F. Cárcano, O.B., B.S., K.G.N., G.M., D.S., W.R.D., M.A.A.K.F., A.v.B., F.N., J.N.H., N.R., W.Y.H., W.M.L., A. Lori, M.F., M.Z.-M., S.V.S., W.J.M., Biobank Japan Project, A.V., R.D., F. Carusso, L.S.G., K.A., M.A.B., C.A., I.P., S. Ricard, FinnGen, G.S., R.E.B., N.S.V., N.S., G.D.S., A.A., S.B., D.H., N.G., P.P., M.S., A.P., F.I.N., M.J.F., X.Z., L.J.M., M.K., T.E., S.A.C., D.C.C., R.G.U., D.Z., A.M., I.H., A.H., L.F., V. Janout, D.M., V. Jinga, S. Rascu, M.M., S.S., S.M., V.G., B.A.-A., J.M., M.J., L.P., L.H., J. Li, I.L., S.M.B., A.G.S., C.T.G.S., R.B.R., F.P.G., M.D.C., M.P., G.-S.M.L., M.L.F., A.J., S.E.G., A.S., R.H.T., V.S., D.D.T., C.T.B., D.A., E.T.L., W.C.N., V.A.M., A.V.P., J.-C.B., N.D.F., P. Bigot, R.M.R., L.M.C., A.F., B.J.M., C.T., T.K.C., D.M.C., R.H., J.E.E.-P., P.H.A., A.G., P. Brennan and J.G. contributed samples and/or data. All authors critically reviewed the paper. The following authors contributed equally as co-first authors: M.P.P., D.D., M.J.M. and B.R.G. The following authors contributed equally to the work as co-second authors: T.W., D.O., S.C., A.F.I., P.A.S., A. Liu, C.W., S.O.A., J. Larkin, S.C.Z., M.S., K.H., A.H., K.A.L., F. Cárcano, O.B., B.S., K.G.N., G.M., D.S., W.R.D., M.A.A.K.F., A.v.B. and F.N. The following authors contributed equally as co-second-last authors: D.A., E.T.L., W.C.N., V.A.M., A.V.P., L.M.C., J.C.B., N.F., P. Bigot, R.M.R., A.F., B.J.M., C.T., T.K.C., D.M., R.H., J.E.E.-P., P.H.A. and A.G. The following authors jointly supervised this work: P. Brennan, J.G. and S.J.C.
Corresponding authors
Ethics declarations
Competing interests
D.S. has received funds from Janssen for consulting outside of the submitted work. N.S.V. has received grants, personal fees and non-financial support from Bristol Myers Squibb; personal fees and non-financial support from Ipsen and EUSA Pharma; and personal fees from Merck Serono, Pfizer, Eisai Ltd and 4D Pharma, all outside the submitted work. G.D.S. has received educational grants from Pfizer and AstraZeneca; consultancy fees from Pfizer, Merck, EUSA Pharma and MSD; travel expenses from Pfizer and speaker fees from Pfizer, all outside the submitted work. M.S. has received honoraria from Covidien/Medtronic for teaching on courses and speaker fees from Pfizer, all outside the submitted work. L.M.C. has received research funding from BMS, Novartis and GSK, all outside the submitted work. B.J.M. has received funding from the NCCN Hereditary Kidney Cancer Panel and Merck, all outside the submitted work. T.K.C. has received funding from Alkermes, AstraZeneca, Aravive, Aveo, Bayer, Bristol Myers Squibb, Calithera, Circle Pharma, Deciphera Pharmaceuticals, Eisai, EMD Serono, Exelixis, GlaxoSmithKline, Gilead, IQVIA, Infinity, Ipsen, Jansen, Kanaph, Lilly, Merck, Nikang, Nuscan, Novartis, Oncohost, Pfizer, Roche, Sanofi/Aventis, Scholar Rock, Surface Oncology, Takeda, Tempest, Up-To-Date, CME events (Peerview, OncLive, MJH, CCO and others), outside the submitted work. The other authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Christopher Amos and A. Ari Hakimi for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Pleiotropic effects of kidney cancer loci for other cancers and risk factors.
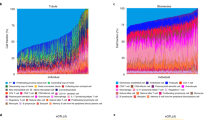
Pleiotropy matrix summarizing kidney cancer susceptibility loci with evidence of pleiotropic effects for other cancers and/or selected risk factors (body mass index, hypertension, blood pressure, smoking) from searches of GWAS Catalog and UK Biobank GWAS summary statistics. Cell colors indicating associations with specific traits: red, other cancers; blue, body mass index; orange, hypertension or blood pressure; violet, smoking. Locus-trait summary statistics listed in Supplementary Table 11 (GWAS Catalog) and 12 (UK Biobank).
Extended Data Fig. 2 In silico analysis of enrichment for putative regulatory annotations among kidney cancer loci.
Enrichment of variants associated with overall RCC in (a) DNAse Hotspots (b) Histone modification sites (c) Chromatin states (d) different genic locations. The enrichments are depicted for variants at different p-value thresholds denoted by the colors and across different categories in each panel. The results were computed using GARFIELD v2.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Supplementary Tables
Supplementary Tables 1–21.
Rights and permissions
About this article
Cite this article
Purdue, M.P., Dutta, D., Machiela, M.J. et al. Multi-ancestry genome-wide association study of kidney cancer identifies 63 susceptibility regions. Nat Genet 56, 809–818 (2024). https://doi.org/10.1038/s41588-024-01725-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-024-01725-7